Metagenome-Atlas
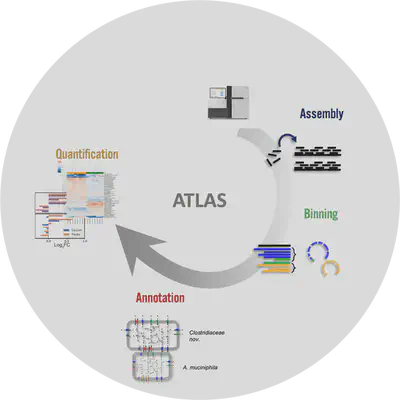
Metagenome-atlas is a easy-to-use metagenomic pipeline. It handles all steps from QC, Assembly, Binning, to Annotation & Quantification.
The pipline is easy to use. You can start using atlas with three commands:
mamba install -y -c bioconda -c conda-forge metagenome-atlas
atlas init --db-dir databases path/to/fastq/files
atlas run all

Atlas has only one dependency: conda; all databases and other dependencies are installed on the fly. Atlas is based on snakemake which enables it to run steps of the workflow in parallel on a cluster.